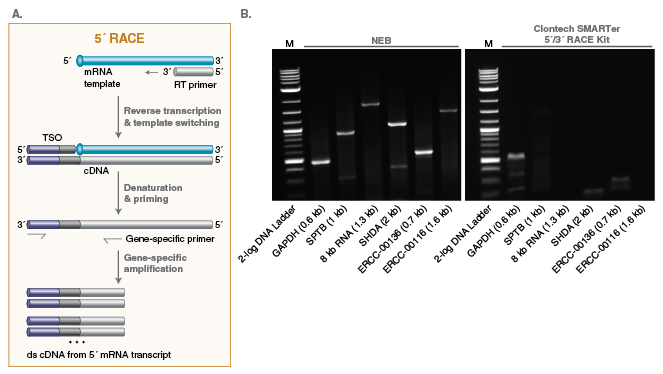
MiR-RACE, a New Efficient Approach to Determine the Precise Sequences of Computationally Identified Trifoliate Orange (Poncirus trifoliata) MicroRNAs | PLOS ONE

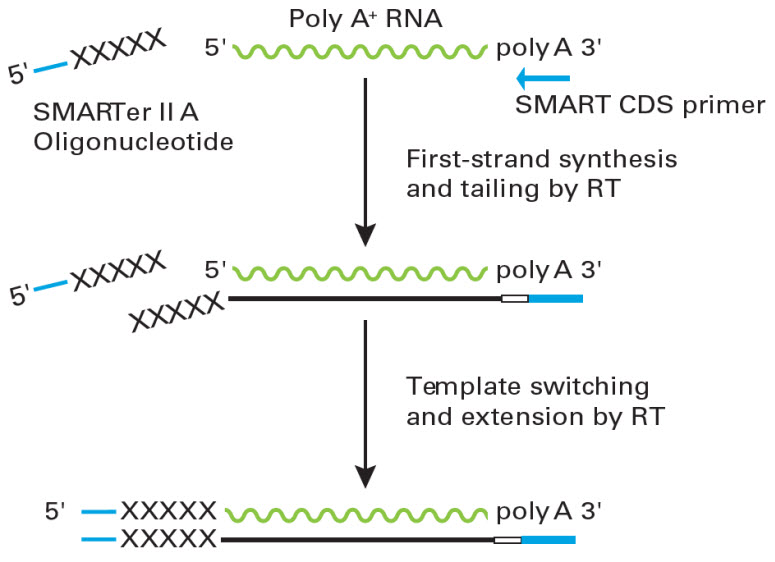
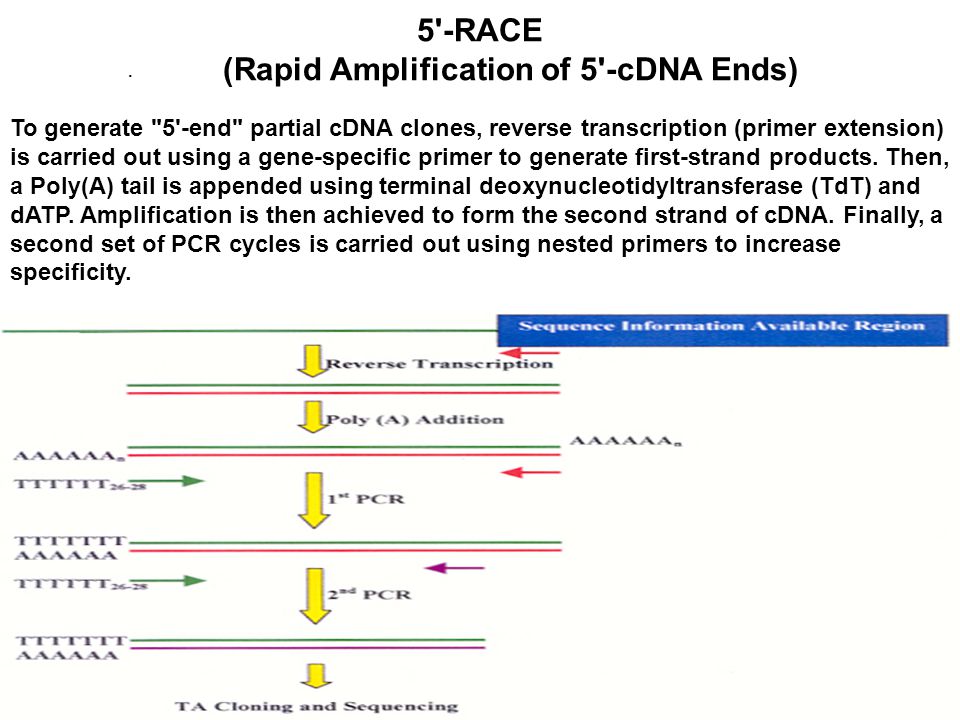
A new rapid amplification of cDNA ends method for extremely guanine plus cytosine-rich genes. - Abstract - Europe PMC
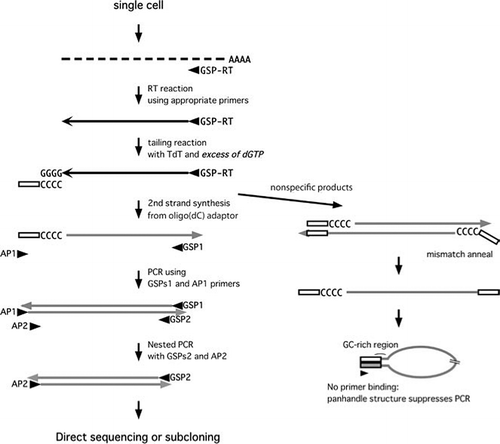
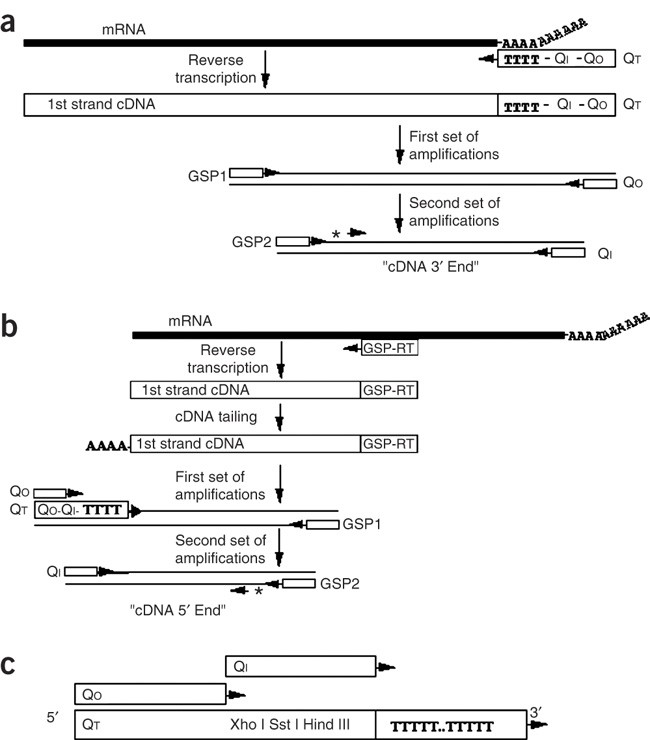
Amplification and analysis of cDNA generated from a single cell by 5′-RACE: application to isolation of antibody heavy and light chain variable gene sequences from single B cells | BioTechniques
Mouse papillomavirus type 1 (MmuPV1) DNA is frequently integrated in benign tumors by microhomology-mediated end-joining | PLOS Pathogens

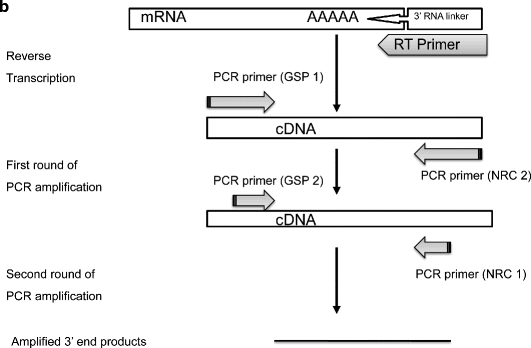
Figure 2 from Targeted rapid amplification of cDNA ends (T-RACE)—an improved RACE reaction through degradation of non-target sequences | Semantic Scholar
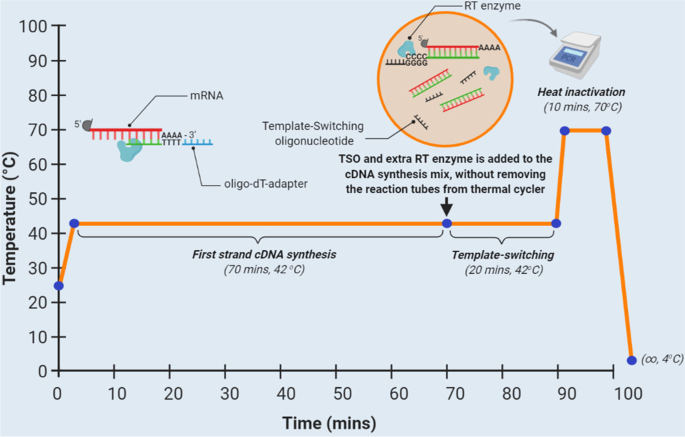
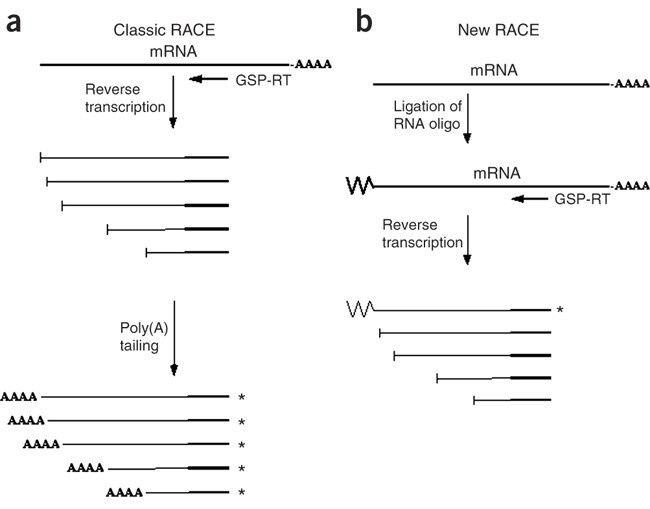
A versatile 5′ RACE-Seq methodology for the accurate identification of the 5′ termini of mRNAs | BMC Genomics | Full Text
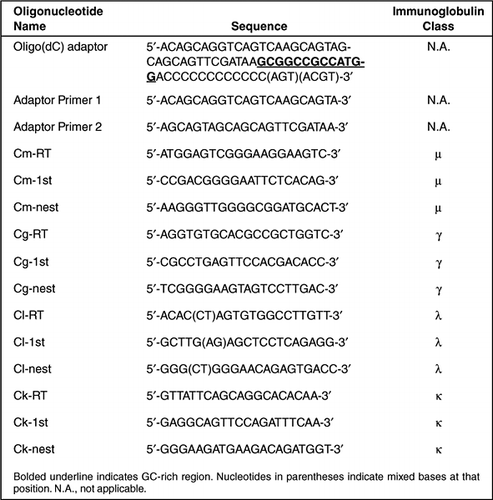
Amplification and analysis of cDNA generated from a single cell by 5′-RACE: application to isolation of antibody heavy and light chain variable gene sequences from single B cells | BioTechniques